Sessions: Interactive work on compute nodes
Questions
How to reach the calculation nodes
How do I proceed to work interactively?
Objectives
be able to start interactive sessions
Be able to run Julia in Jupyter notebook
Compute allocations in this workshop
Rackham:
naiss2024-22-1202Kebnekaise:
hpc2n2024-114Cosmos:
lu2024-7-80
Storage space for this workshop
Rackham:
/proj/r-py-jl-m-rackhamKebnekaise:
/proj/nobackup/r-py-jl-mCosmos:
<your own good place>
Note
It is possible to run Julia directly on the login (including ThinLinc) nodes.
But this should only be done for shorter jobs or jobs that do not use a lot of resources, as the login nodes can otherwise become slow for all users.
If you want to work interactively with your code or data, you should start an interactive session.
If you rather will run a script which won’t use any interactive user input while running, you can instead start a batch job, see next session.
There are several ways to run Julia interactively
Directly on the login nodes: only do this for short jobs that do not take a lot of resources
As an interactive job on the computer nodes, launched via the batch system
Jupyter notebooks on compute node.
General
In order to run interactively, you need to have compute nodes allocated to run on, and this is done through the Slurm system.
Because you will have to wait until the nodes are allocated, and because you cannot know when this happens, this is not usually a recommended way to run Julia, but it is possible.
Warning
(HPC2N) Do note that it is not real interactivity as you probably mean it, as you will have to run it as a Julia script instead of by starting Julia and giving commands inside it.
The reason for this is that you are not actually logged into the compute node and only sees the output of the commands you run.
Julia “interactively” on the compute nodes
Note
- On UPPMAX and LUNARC:
interactive ... You get graphics as well!
- On UPPMAX and LUNARC:
- On HPC2N:
salloc This command works as well on the other clusters but brings no or bad graphics.
- On HPC2N:
When the resources are allocated, you need to preface commands with
srunin order to run on the allocated nodes instead of the login node.
First, you make a request for resources with
interactive/salloc, like this:
$ interactive -n <tasks> --time=HHH:MM:SS -A naiss2024-22-1202
$ salloc -n <tasks> --time=HHH:MM:SS -A hpc2n2023-114
$ interactive -n <tasks> --time=HHH:MM:SS -A lu2024-7-80
where <tasks> is the number of tasks (or cores, for default 1 task per core), time is given in hours, minutes, and seconds (maximum T168 hours), and then you give the id for your project
Your request enters the job queue just like any other job, and interactive/salloc will tell you that it is waiting for the requested resources.
When salloc tells you that your job has been allocated resources, you can interactively run programs on those resources with
srun.The commands you run with
srunwill then be executed on the resources your job has been allocated.
On HPC2N
If you do not preface with
srunthe command is run on the login node!You can now run Julia scripts on the allocated resources directly instead of waiting for your batch job to return a result.
This is an advantage if you want to test your Julia script or perhaps figure out which parameters are best.
Documentation at the centers
Example Code along
Type-Along
Requesting 4 cores for 10 minutes, then running Julia
[bjornc@rackham2 ~]$ interactive -A naiss2024-22-1202 -p core -n 4 -t 10:00
You receive the high interactive priority.
There are free cores, so your job is expected to start at once.
Please, use no more than 6.4 GB of RAM.
Waiting for job 29556505 to start...
Starting job now -- you waited for 1 second.
[bjornc@r483 ~]$ module load julia/1.8.5
Let us check that we actually run on the compute node:
[bjornc@r483 ~]$ srun hostname
r483.uppmax.uu.se
r483.uppmax.uu.se
r483.uppmax.uu.se
r483.uppmax.uu.se
We are. Notice that we got a response from all four cores we have allocated.
[~]$ salloc -n 4 --time=00:30:00 -A hpc2n2024-114
salloc: Pending job allocation 20174806
salloc: job 20174806 queued and waiting for resources
salloc: job 20174806 has been allocated resources
salloc: Granted job allocation 20174806
salloc: Waiting for resource configuration
salloc: Nodes b-cn0241 are ready for job
[~]$ module load GCC/11.2.0 OpenMPI/4.1.1 julia/1.8.5
[~]$
Let us check that we actually run on the compute node:
[~]$ srun hostname
b-cn0241.hpc2n.umu.se
b-cn0241.hpc2n.umu.se
b-cn0241.hpc2n.umu.se
b-cn0241.hpc2n.umu.se
We are. Notice that we got a response from all four cores we have allocated.
[bjornc@cosmos1 ~]$ interactive -A lu2024-7-80 -n 4 -t 10:00
Cluster name: COSMOS
Waiting for JOBID 930844 to start
[bjornc@cn050 ~]$ module load Julia/1.8.5-linux-x86_64
Let us check that we actually run on the compute node:
[bjornc@cn050 ~]$ echo $SLURM_CPUS_ON_NODE
4
We are, because the $SLURM* environment variable gves an output. Notice that we got 4, whihc is nt the size of the physcial node bt the allocation size.
Running a script
- The script
Adding two numbers from user input (serial-sum.jl)
# This program will add two numbers that are provided by the user # Get the numbers x = parse( Int32, ARGS[1] ) y = parse( Int32, ARGS[2] ) # Add the two numbers together summ = x + y println("The sum of the two numbers is ", summ)
Running the script
Note that the commands are the same for both HPC2N and UPPMAX!
Running a Julia script in the allocation we made further up. Notice that since we asked for 4 cores, the script is run 4 times, since it is a serial script
[~]$ srun julia serial-sum.jl 3 4 The sum of the two numbers is: 7 The sum of the two numbers is: 7 The sum of the two numbers is: 7 The sum of the two numbers is: 7 [~]$
Without the
sruncommand, Julia won’t understand that it can use several cores. Therefore the program is run only once.[~]$ julia serial-sum.jl 3 4 The sum of the two numbers is: 7
Running Julia REPL (UPPMAX/HPC2N)
First start Julia using the 4 cores and check if workers are available
$ julia -p 4
julia> nworkers()
4
Exit
When you have finished using the allocation, either wait for it to end, or close it with exit
[bjornc@r483 ~]$ exit
exit
[screen is terminating]
Connection to r483 closed.
[bjornc@rackham2 ~]$
[~]$ exit
exit
salloc: Relinquishing job allocation 20174806
salloc: Job allocation 20174806 has been revoked.
[~]$
[~]$ exit
exit
[screen is terminating]
Connection to cn050 closed.
[~]$
Running IJulia and Jupyter
For more interactiveness you can run IJulia.
You benefit a lot if you are using ThinLinc
Like for Python it is possible to run Julia in Jupyter, i.e. in a web interface with possibility of inline figures and debugging. An easy way to do this is to load the python module as well. In shell:
$ module load julia/1.8.5 $ module load python/3.9.5 $ julia -p 4
In Julia:
julia> using Pkg julia> Pkg.add("IJulia") julia> Pkg.build("IJulia") julia> using IJulia julia> notebook(dir=".",detached=true)
A Firefox session should start with the Jupyter notebook interface.
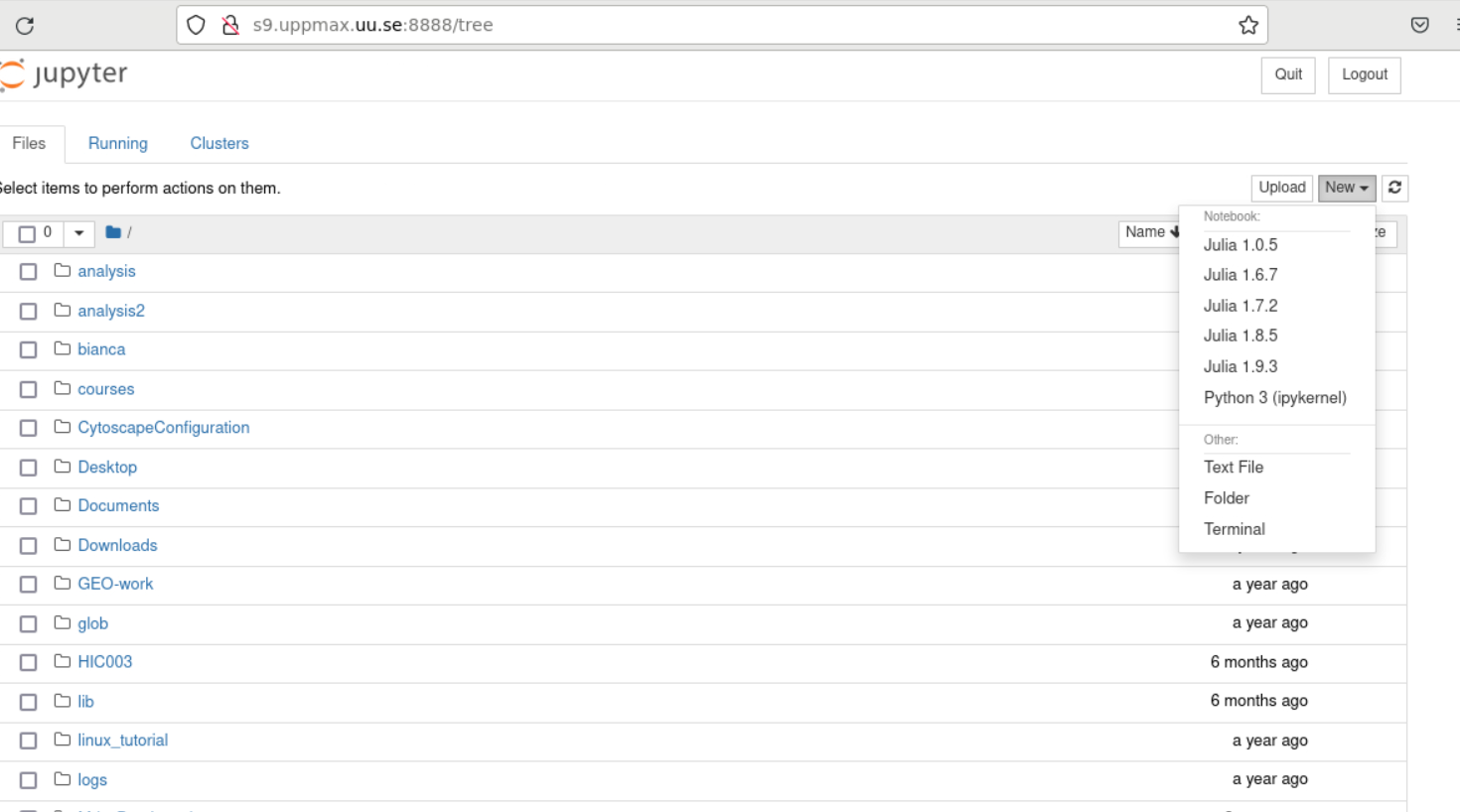
Note
You only have to add and build IJulia the first time for each julia version and each jupyter, provided with a python version at UPPMAX
Tip
With notebook(dir="</path/to/work/dir/>", detached=true) the notebook will not be killed when you exit your REPL Julia session in the terminal.
Running IJulia in Jupyter on compute nodes
Jupyter is rather slow on the compute nodes.
This can be fixed by opening jupyter in a web browsers on your local computer or in ThinLinc
Remember to load python as well and to go via the
julia -p <number of cores>andnotebook(<options>)inside the Julia session instead of startingjupiter-notebookin the bash shell.
Jupyter from terminal
If IJulia is precompiled once then you can run Julia from Jupyter directly from the terminal
Start the browser from the ThinLinc menu.
Copy-paste one of the address lines from the jupyter output
You can start the Julia kernel in the upper right corner!
Like for Python it is possible to run a Julia in a Jupyter, i.e. in a web interface with possibility of inline figures and debugging. An easy way to do this is to load the JupyterLab and Julia modules. In shell:
$ module load GCCcore/13.2.0 JupyterLab/4.2.0
$ module load Julia/1.8.5-linux-x86_64
$ julia
In Julia package mode:
(v1.8) pkg>add IJulia
(v1.8) pkg>build IJulia
Write a bash script similar to this (call it job_jupyter.sh, for instance):
#!/bin/bash
# Here you should put your own project id
#SBATCH -A hpc2n2024-114
# This example asks for 1 core
#SBATCH -n 1
# Ask for a suitable amount of time. Remember, this is the time the Jupyter notebook will be available! HHH:MM:SS.
#SBATCH --time=00:10:00
# Clear the environment from any previously loaded modules
module purge > /dev/null 2>&1
# Load the module environment suitable for the job
module load GCCcore/13.2.0 JupyterLab/4.2.0
# Load the Julia module
ml Julia/1.8.5-linux-x86_64
# Start JupyterLab
jupyter lab --no-browser --ip $(hostname)
Then, in the output file slurm-<jobID>.out file, copy the url that starts with http://b-cn1403.hpc2n.umu.se:8888/lab and paste it in a Firefox browser on Kebnekaise. When the Jupyter notebook interface starts, you can choose the Julia version from the module you loaded (in this case 1.8.5).
Running Julia in Jupyter on compute nodes at HPC2N
On Kebnekaise, you can run Jupyter notebooks with Julia kernels by using batch scripts
https://docs.hpc2n.umu.se/tutorials/jupyter/#jupyterlab__with__julia
Exercises
1. Try to run scripts from an interactive session
Try out one or two of the scripts from the exercise folder
batchJulia.- First create an interactive session with the right Slurm commands to the
interactive/salloccommand. use the commands from the batch job script belonging to the julia script at examples of batch scripts for julia
- First create an interactive session with the right Slurm commands to the
Keypoints
Start an interactive session on a calculation node by a SLURM allocation
At HPC2N:
salloc…At UPPMAX/LUNARC:
interactive…
Follow the same procedure as usual by loading the Julia module and possible prerequisites.
Run Julia in Jupyter lab/notebook
Procedure is to use the IJulia package and start a jupyter notebook from the julia command line.